Search
- Page Path
- HOME > Search
Original Articles
- The first reported hepatitis E outbreak in a food manufacturing factory: Korea, 2022
- Hansol Yeom, Soonryu Seo, Youngsil Yoon, Jaeeun Lee, Myung-Guk Han, Deog-Yong Lee, Sun-Whan Park, Song A Park, Sook-Hyang Jeong, Jin Gwack
- Osong Public Health Res Perspect. 2023;14(1):15-22. Published online February 22, 2023
- DOI: https://doi.org/10.24171/j.phrp.2022.0305
- 1,663 View
- 139 Download
-
 Graphical Abstract
Graphical Abstract
 Abstract
Abstract
 PDF
PDF 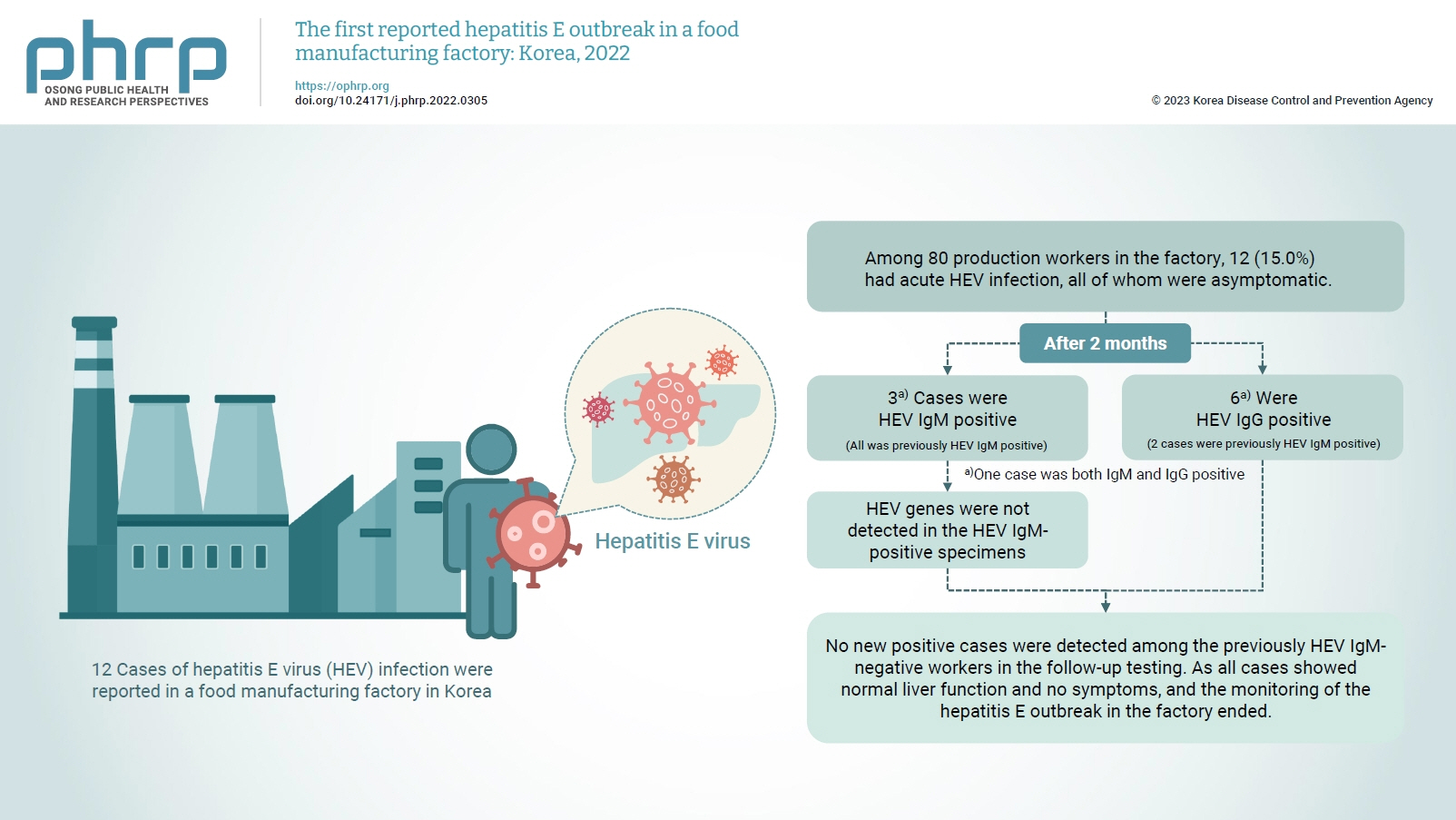
- Objectives
On February 16, 2022, 12 cases of hepatitis E virus (HEV) infection were reported in a food manufacturing factory in Korea. The aim of this study was to identify additional cases and to determine the source of this HEV outbreak. Methods: This study was an in-depth investigation of 12 HEV immunoglobulin M (IgM)-positive cases and their demographic, clinical, and epidemiological characteristics. On-site specimens were collected from the environment and from humans, and a follow-up investigation was conducted 2 to 3 months after the outbreak. Results: Among 80 production workers in the factory, 12 (15.0%) had acute HEV infection, all of whom were asymptomatic. The follow-up investigation showed that 3 cases were HEV IgMpositive, while 6 were HEV IgG-positive. HEV genes were not detected in the HEV IgM-positive specimens. HEV genes were not detected in the food products or environmental specimens collected on-site. HEV was presumed to be the causative pathogen. However, it could not be confirmed that the source of infection was common consumption inside the factory. Conclusion: This was the first domestic case of an HEV infection outbreak in a food manufacturing factory in Korea. Our results provide information for the future control of outbreaks and for the preparation of measures to prevent domestic outbreaks of HEV infection.
- One-Step Reverse Transcription-Polymerase Chain Reaction for Ebola and Marburg Viruses
- Sun-Whan Park, Ye-Ji Lee, Won-Ja Lee, Youngmee Jee, WooYoung Choi
- Osong Public Health Res Perspect. 2016;7(3):205-209. Published online June 30, 2016
- DOI: https://doi.org/10.1016/j.phrp.2016.04.004
- 2,817 View
- 23 Download
- 5 Crossref
-
 Abstract
Abstract
 PDF
PDF - Objectives
Ebola and Marburg viruses (EBOVs and MARVs, respectively) are causative agents of severe hemorrhagic fever with high mortality rates in humans and nonhuman primates. In 2014, there was a major Ebola outbreak in various countries in West Africa, including Guinea, Liberia, Republic of Sierra Leone, and Nigeria. EBOV and MARV are clinically difficult to diagnose and distinguish from other African epidemic diseases. Therefore, in this study, we aimed to develop a method for rapid identification of the virus to prevent the spread of infection.
Methods
We established a conventional one-step reverse transcription-polymerase chain reaction (RT-PCR) assay for these pathogens based on the Superscript Reverse Transcriptase-Platinum Taq polymerase enzyme mixture. All assays were thoroughly optimized using in vitro-transcribed RNA.
Results
We designed seven primer sets of nucleocapsid protein (NP) genes based on sequences from seven filoviruses, including five EBOVs and two MARVs. To evaluate the sensitivity of the RT-PCR assay for each filovirus, 10-fold serial dilutions of synthetic viral RNA transcripts of EBOV or MARV NP genes were used to assess detection limits of viral RNA copies. The potential for these primers to cross react with other filoviruses was also examined. The results showed that the primers were specific for individual genotype detection in the examined filoviruses.
Conclusion
The assay established in this study may facilitate rapid, reliable laboratory diagnosis in suspected cases of Ebola and Marburg hemorrhagic fevers. -
Citations
Citations to this article as recorded by- Marburg Virus Disease – A Mini-Review
Sandip Chakraborty, Deepak Chandran, Ranjan K. Mohapatra, Mahmoud Alagawany, Mohd Iqbal Yatoo, Md. Aminul Islam, Anil K. Sharma, Kuldeep Dhama
Journal of Experimental Biology and Agricultural S.2022; 10(4): 689. CrossRef - Marburgviruses: An Update
Caterina M Miraglia
Laboratory Medicine.2019; 50(1): 16. CrossRef - Ebola virus: A global public health menace: A narrative review
Shamimul Hasan, SyedAnsar Ahmad, Rahnuma Masood, Shazina Saeed
Journal of Family Medicine and Primary Care.2019; 8(7): 2189. CrossRef - Fast and Parallel Detection of Four Ebola Virus Species on a Microfluidic-Chip-Based Portable Reverse Transcription Loop-Mediated Isothermal Amplification System
Xue Lin, Xiangyu Jin, Bin Xu, Ruliang Wang, Rongxin Fu, Ya Su, Kai Jiang, Han Yang, Ying Lu, Yong Guo, Guoliang Huang
Micromachines.2019; 10(11): 777. CrossRef - The current landscape of nucleic acid tests for filovirus detection
David J. Clark, John Tyson, Andrew D. Sails, Sanjeev Krishna, Henry M. Staines
Journal of Clinical Virology.2018; 103: 27. CrossRef
- Marburg Virus Disease – A Mini-Review



 First
First Prev
Prev


